
鄭小琪,博士,教授
上海交通大學(xué)公共衛(wèi)生學(xué)院,
單細(xì)胞組學(xué)與疾病研究中心,生物信息學(xué)平臺(tái)主任。
通訊地址:上海市黃浦區(qū)重慶南路227號(hào)
郵箱: [email protected] 或 [email protected]
個(gè)人網(wǎng)站: https://xiaoqizheng.github.io/index.html
個(gè)人簡(jiǎn)介:
2009年畢業(yè)于大連理工大學(xué)應(yīng)用數(shù)學(xué)系獲理學(xué)博士學(xué)位,隨后在上海師范大學(xué)數(shù)學(xué)系工作,于 2017年晉升至教授、博士生導(dǎo)師。2022年底,加入上海交通大學(xué)公共衛(wèi)生學(xué)院,任單細(xì)胞組學(xué)與疾病研究中心生物信息學(xué)平臺(tái)主任。
主要從事生物統(tǒng)計(jì)和生物信息學(xué)領(lǐng)域的研究,擅長(zhǎng)生物醫(yī)學(xué)大數(shù)據(jù)的統(tǒng)計(jì)建模與算法開(kāi)發(fā),機(jī)器學(xué)習(xí)、深度學(xué)習(xí)等人工智能方法在數(shù)據(jù)挖掘中的應(yīng)用等。在基于DNA甲基化 (包括亞硫酸鹽測(cè)序和450k芯片數(shù)據(jù)) 的腫瘤純度估計(jì)、校正腫瘤純度的差異甲基化分析及腫瘤樣本聚類等問(wèn)題上取得了一系列研究成果。2008年至今累計(jì)發(fā)表SCI論文60余篇,包括第一或通訊作者論文40余篇,累積影響因子超過(guò)300。主持上海市人才發(fā)展基金,以負(fù)責(zé)人承擔(dān)國(guó)家自然科學(xué)基金面上項(xiàng)目2項(xiàng),青年項(xiàng)目1項(xiàng),以項(xiàng)目骨干承擔(dān)國(guó)家重點(diǎn)研發(fā)計(jì)劃專項(xiàng)1項(xiàng)。現(xiàn)任中國(guó)工業(yè)與應(yīng)用數(shù)學(xué)學(xué)會(huì)(CSIAM)數(shù)學(xué)生命科學(xué)分會(huì)常務(wù)理事,上海非線性科學(xué)研究會(huì)副秘書(shū)長(zhǎng),中國(guó)計(jì)算機(jī)學(xué)會(huì)生物信息專業(yè)委員會(huì)委員等。
研究方向:
1. 單細(xì)胞轉(zhuǎn)錄組與空間轉(zhuǎn)錄組數(shù)據(jù)分析
2. 基于DNA甲基化的腫瘤異質(zhì)性分析
3. 生物醫(yī)學(xué)研究中的人工智能方法(包括統(tǒng)計(jì)學(xué)習(xí)和深度學(xué)習(xí))
教育經(jīng)歷:
2004 - 2009, 博士,應(yīng)用數(shù)學(xué), 大連理工大學(xué)
2000 - 2004, 學(xué)士,數(shù)學(xué)與應(yīng)用數(shù)學(xué),山東師范大學(xué)
工作經(jīng)歷:
2022.11 – 至今, 教授, 上海交通大學(xué)公共衛(wèi)生學(xué)院,單細(xì)胞組學(xué)與疾病研究中心
2017.09 - 2022.11, 教授, 上海師范大學(xué)數(shù)學(xué)系
2018.07 - 2018.09, 訪問(wèn)學(xué)者, 埃默里大學(xué)公共衛(wèi)生學(xué)院,生物統(tǒng)計(jì)與生物信息學(xué)系
2012.09 - 2017.09, 副教授, 上海師范大學(xué)數(shù)學(xué)系.
2012.12 - 2014.02, 訪問(wèn)學(xué)者, 哈佛大學(xué)公共衛(wèi)生學(xué)院,Dana-Farber癌癥研究中心
2009.06 - 2012.09, 講師, 上海師范大學(xué)數(shù)學(xué)系
代表性論文 (#: 共同第一, *: 共同通訊) :
[1] Wei N, Nie Y, Liu L*, Zheng X*, Wu HJ*, Secuer: Ultrafast, scalable and accurate clustering of single-cell RNA-seq data, PLoS Computational Biology 2022, doi.org/10.1371/journal.pcbi.1010753.
[2] Ding Y, Cai K, Liu L, Zhang Z, Zheng X*, Shi J*, mHapTk: a comprehensive toolkit for the analysis of DNA methylation haplotypes, Bioinformatics. 2022, 38(22):5141-5143.
[3] Zhang Z#, Dan Y#, Xu Y, Zhang J, Zheng X*, Shi J*: The DNA methylation haplotype (mHap) format and mHapTools, Bioinformatics 2021, 37 (24):4892-4894.
[4] Zhang W#, Li Z, Wei N, Wu H-J, Zheng X*: Detection of differentially methylated CpG sites between tumor samples with uneven tumor purities. Bioinformatics 2020, 36(7):2017-2024.
[5] Qin Y#, Zhang W#, Sun X, Nan S, Wei N, Wu HJ, Zheng X*: Deconvolution of heterogeneous tumor samples using partial reference signals. PLoS Computational Biology 2020, 16(11): e1008452.
[6] T Xu, X Zheng, B Li, P Jin, Z Qin, H Wu: A comprehensive review of computational prediction of genome-wide features, Briefings in bioinformatics 2020, 21(1), 120-134.
[7] Liu X, Liu T, Shang Y, Dai P, Zhang W, Lee BJ, Huang M, Yang D, Wu Q, Liu LD, Zheng X, Zhou BO, Dong J, Yeap LS, Hu J, Xiao T, Zha S, Casellas R, Liu XS, Meng FL: ERCC6L2 promotes DNA orientation-specific recombination in mammalian cells, Cell research 2020, 30 (9):732-744.
[8] Hu X, Zhang J, Wang J, Fu J, Li T, Zheng X, Wang B, Gu S, Jiang P, Fan J, Carroll, MC, Wucherpfennig K, Hacohen N, Zhang F, Zhang P, Liu JS*, Li B*, Liu XS*: Widespread B cell clonal expansions and related mechanism of immune evasion in human cancers, Nature Genetics 2019, 51(3):560-567.
[9] Fan H#, Lv P#, Huo X, Wu J, Wang Q, Cheng L, Liu Y, Tang Q, Zhang L, Zhang F, Zheng X, Wu H, Wen B*: The nuclear matrix protein HNRNPU maintains 3D genome architecture globally in mouse hepatocytes, Genome Research, 2018, 28:192-202.
[10] Niu B#, Paulson JN, Zheng X, Kolter R*: Simplified and Representative Bacterial Community of Maize Roots. P Natl Acad Sci USA 2017, 114 (12): E2450-E2459.
[11] Zheng X#*, Zhang N#, Wu HJ, Wu H*: Estimating and accounting for tumor purity in the analysis of DNA methylation data from cancer studies. Genome Biology 2017, 18:17.
[12] Zheng X#*, Zhong S: From structure to function, how bioinformatics help to reveal functions of our genomes. Genome Biology 2017, 18:183.
[13] Zhang W#, Feng H#, Wu H*, Zheng X*, Accounting for Tumor purity improves cancer subtype classification from DNA methylation data, Bioinformatics 33(17), 2017, 2651–2657.
[14] Ni T#, Li XY#, Lu N#, An T, Liu ZP, Fu R, Lv WC, Zhang YW, Xu XJ, Grant Rowe R, Lin YS, Scherer A, Feinberg T, Zheng XQ, Chen BA, Liu XS, Guo QL, Wu ZQ*, Weiss SJ*: Snail1-dependent p53 repression regulates expansion and activity of tumour-initiating cells in breast cancer. Nat Cell Biol 2016, 18:1221-1232.
[15] Wang F#, Zhang N, Wang J, Wu H#, Zheng X*: Tumor purity and differential methylation in cancer epigenomics. Briefings in Functional Genomics 2016, 15:408-419.
[16] Zhang N#, Wu HJ#, Zhang W, Wang J, Wu H*, Zheng X*: Predicting tumor purity from methylation microarray data. Bioinformatics 2015, 31:3401-3405.
[17] Zhang N#, Wang H#, Fang Y, Wang J*, Zheng X*, Liu XS*: Predicting Anticancer Drug Responses Using a Dual-Layer Integrated Cell Line-Drug Network Model. PLoS Computational Biology 2015, 11:e1004498.
[18] Dong Z#, Zhang N#, Li C, Wang H, Fang Y, Wang J, Zheng X*: Anticancer drug sensitivity prediction in cell lines from baseline gene expression through recursive feature selection. BMC Cancer 2015, 15:489.
[19] Zheng X#, Zhao Q#, Wu H-J#, Li W, Wang H, Meyer CA, Qin QA, Xu H, Zang C, Jiang P, Li F, Hou Y, He J, Wang J, Wang J, Zhang P*, Zhang Y*, Liu XS*: MethylPurify: tumor purity deconvolution and differential methylation detection from single tumor DNA methylomes. Genome Biology 2014, 15:419.
[20] Li L#, Yu S, Xiao W, Li Y, Huang L, Zheng X*, Zhou S*, Yang H*: Sequence-based identification of recombination spots using pseudo nucleic acid representation and recursive feature extraction by linear kernel SVM. BMC bioinformatics 2014, 15:340.
[21] Zhu J#, Qin Y, Liu T, Wang J, Zheng X*: Prioritization of candidate disease genes by topological similarity between disease and protein diffusion profiles. BMC bioinformatics 2013, 14:S5.
[22] Wang J#, Zheng X*: Comparison of protein secondary structures based on backbone dihedral angles. J Theor Biol 2008, 250:382-387.
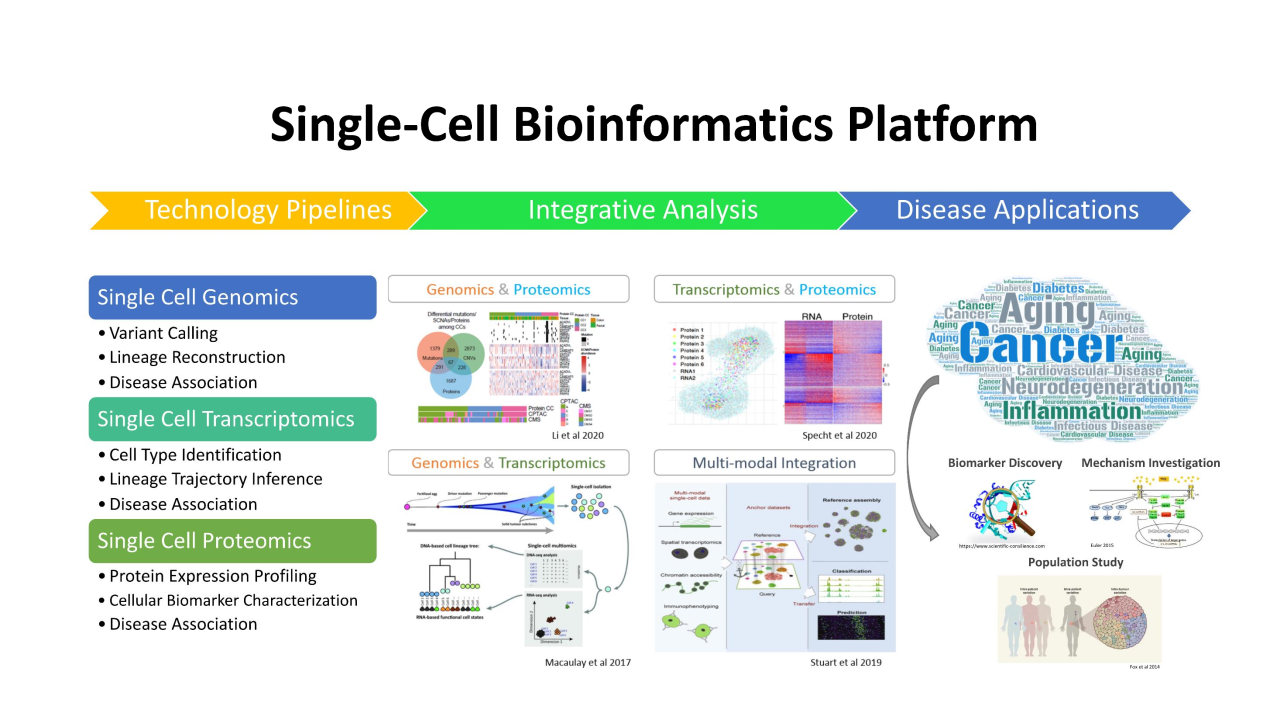
單細(xì)胞生物信息學(xué)平臺(tái)
平臺(tái)主任:鄭小琪
技術(shù)員:胥宛星